PRKN Variant Classification Criteria And Portugal ARJP Mutation Summary
PRKN Variant Annotation Continuation
Variant fragments:
RORB domain
p.T240R
p.R275W (= p.Arg275Tryptophan) = R>W, C.823>T, or rs34424986, VCV000007050
R275Q: p.Arg275Gln
c.824G>A Variation ID: 931482, Accession: VCV000931482.2
p.G430D (= p.Gly430Asp)
p.K161N
P.c253FAuthor note:
이건 원문에 없으나 내가 clinvar 로부터 추가
rs 번호는 없네
Js: these two were not found as clinical pathogenic, but from functional assays (mitophagy, protein expression), these two look pathogenicNote attached to p.R275W:
FATHMM prediction Pathogenic score: score 0.87 (이건 cancer 용인가?) is with
https://cancer.sanger.ac.uk/cosmic/mutation/overview/id=98884592
Yi 2019 의 suppl table 보면 (excel), benign point from MAF in population 이 1임,
이건 normal people 에서도 일부 발견된다는 말 아닌가?
그러나 Benign points from homozygotes in population 가 0인걸 보면,
적어도 homoz는 일반에서 안 발견되니 봄p.G430D is visually associated with (Mortiboys, 2008 #719).
ACMG-AMP / Yi Pathogenic vs Benign Criteria
Note) [methods] 이 전체가 ACMG-AMP criteria
1. Clinical evidence (from public database) 먼저:
Pathogenic vs Benign (scoring system, algorithm-based) (2019 Yi)Criteria summary:
| side | criteria / notes |
|---|---|
| Pathogenic | the more patients in the family homoz or compound heteroz in Parkin ('Segregation within family', the more pathogenic |
| Pathogenic note | suppl table 보면 (excel), 13개 pathogenic variants 는 모두 Benign points from homozygotes in population 가 0임. 즉 거꾸로 0인 것들만 모아서 pathogic 이라고 했다는 뜻. |
| Benign | Age > 50 |
| Benign note | The more common in population database (ExAC, Exosome variant serve, dbSNP), the more benign, (even if they are homoz!) |
이렇게 해서 13 pathogenic & likely pathogenic variants 정리Summary For Clinical Trial From Yi
Title:
Summary for clinical trial from Yi (임상에서 만날 종류들, only missense mutations)| group | Yi category | causality statement | mitophagy | include in clinical trials? |
|---|---|---|---|---|
| All homoz or compound heteroz Parkin mutation | found in PD patients: 75종 (24+51), js: these are the patients we will find in a clinical trial | |||
Pathogenic by Yi’s study: 13 | Definitely parkin mutation caused PD. | All ↓ mitophagy | Include | |
benign by Yi’s study: 10 | Yi: benign means not disease causing | Some ↓, Some =, some ↑ | Exclude because 1. Not disease-causing, 2. No mitophagy reduction | |
Uncertain by Yi’s study: 52 (69%, 52/75) | We are not sure if the mutation caused PD because 1. The mutation is also found in normal people (31 cases), or 2. The mutation is found in Late onset PD (probably 21 cases). | unknown | For this 70% of all mutations, 1. we don’t know if the mutation is the cause of PD, 2. We don’t know mitophagy is reduced or not. -> 확인 필요 | |
| Found only in population | not in PD patients: 164 |
Masten 2018 Systematic Review / MDSGene Frequency
Masten, 2018 #708) all mutations, systematic review
1350 sequence variants reported in index patients
139 disease-causing sequence variants
35 definitely pathogenic
94 probably pathogenic
10 possibly pathogenic
Non-disease causing sequence variants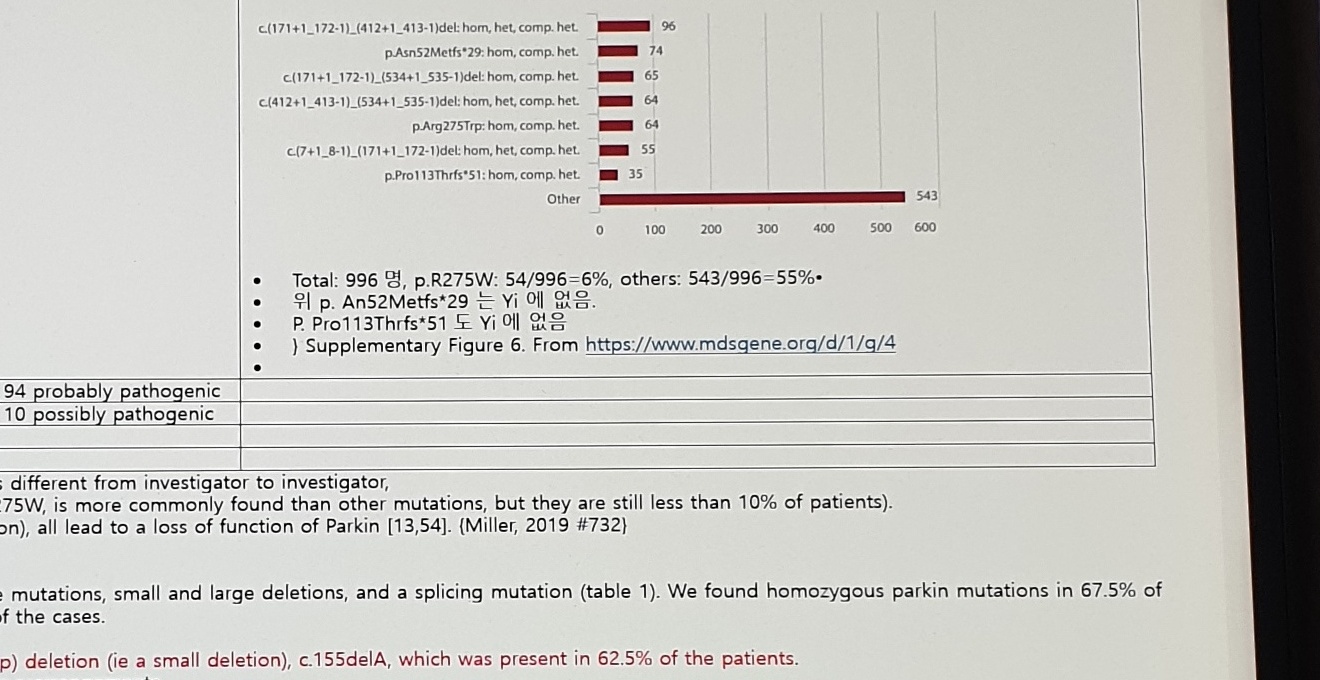
Figure title and note:
Frequency of mutations
Number of patients with missing data: 0 (0.0%)
Supplementary Figure 6. From https://www.mdsgene.org/d/1/g/4Bar-label/value fragments:
| mutation label | count |
|---|---|
c.(171+1_172-1)_(412+1_413-1)del: hom, het, comp. het. | 96 |
p.Asn52Metfs*29: hom, comp. het. | 74 |
c.(171+1_172-1)_(534+1_535-1)del: hom, comp. het. | 65 |
c.(412+1_413-1)_(534+1_535-1)del: hom, het, comp. het. | 64 |
p.Arg275Trp: hom, comp. het. | 64 |
c.(7+1_8-1)_(171+1_172-1)del: hom, het, comp. het. | 55 |
p.Pro113Thrfs*51: hom, comp. het. | 35 |
Other | 543 |
Note below the chart:
Total: 996명, p.R275W: 54/996=6%, others: 543/996=55%
위 p.Asn52Metfs*29 는 Yi 에 없음.
p.Pro113Thrfs*51 도 Yi 에 없음Conclusion text:
Definition of pathogenic (ie 'disease causing') mutation is different from investigator to investigator.
Mutations are variable, although a few mutation, eg p.R275W, is more commonly found than other mutations, but they are still less than 10% of patients).
deletions, truncations and exon duplications (multiplication), all lead to a loss of function of Parkin [13,54]. (Miller, 2019 #732)Morais 2016 Portugal Mutation Summary
Morais, 2016 #914) Portugal
We identified 18 different mutations, including missense mutations, small and large deletions, and a splicing mutation (table 1).
We found homozygous parkin mutations in 67.5% of the patients, and large deletions were present in 42.5% of the cases.
Small deletion
- The most frequent mutation was a 1-base pair (bp) deletion (ie a small deletion), c.155delA, which was present in 62.5% of the patients.
Seventeen patients (js 17/40 = 42.5%) showed large gene rearrangements,
- 9 different deletions either in homozygosity or heterozygosity.
- The most common deletions were those of exon 4 and of exons 3-6 (table 1)
Js: from the below table, 21 patients had a family history, so half!Portugal 40-Patient Molecular Diagnosis Table
Title/header fragments:
Table 1
Overview of molecular and clinical information from 40 patients with a molecular diagnosis of autosomal recessive juvenile Parkinson disease
Patient
Age at onset, y
cDNA
Protein
Family history
ConsanguinityRepeated values:
c.155delA
p.N52Mfs*29
c.823C>T
p.R275W
p.R42P
p.L358Rfs*77
p.R366Q
p.E344Sfs*91
Yes
NoSelected row fragments near the top of the table:
| visible age at onset | visible cDNA/protein fragment | family history | consanguinity |
|---|---|---|---|
22 | c.1244C>A + c.172-52958_734+8943del; [p.T415N] + [p.?] | Yes | No |
28 | c.171+67708_734+58232delins28 repeated; [p.?] + [p.?] | No | No |
38 | c.155delA + c.155delA; [p.N52Mfs*29] + [p.N52Mfs*29] | No | Yes |
25 | c.155delA + c.155delA; [p.N52Mfs*29] + [p.N52Mfs*29] | Yes | No |
13 | c.155delA + c.823C>T; [p.N52Mfs*29] + [p.R275W] | Yes | No |
17 | c.823C>T + c.823C>T; [p.R275W] + [p.R275W] | Yes | No |
33 | c.125G>C + c.125G>C; [p.R42P] + [p.R42P] | No | No |
55 | c.1097G>A + c.413-10504_534+408delins20; [p.R366Q] + [p.?] | No | No |
34 | c.1030delG + c.1286-3C>G; [p.E344Sfs*91] + [p.?] | No | No |
Uncertain Spans
| location | text/status | reason |
|---|---|---|
| page label | Page 71 of 77 | Apple Vision and PaddleOCR agree, but adjacent photos can share the same Word page label during scrolling. |
| nav path | Parkin > Parkin PD | Body content is PRKN/Parkin PD evidence; nav highlight is visible but not a clean single active line. |
| variant table continuation | RORB domain, R275Q, p.K161N, P.c253F, p.G430D, clinical/functional annotation columns | Dense continuation from adjacent photo; row/column alignment remains uncertain. |
p.R275W note | FATHMM, score 0.87, COSMIC URL, Yi benign-point interpretation | Text is small and partly inside a table cell; OCR differs on URL punctuation. |
| Yi criteria table | 13 pathogenic variants, Benign points from homozygotes in population, Age > 50, ExAC, Exosome variant serve, dbSNP | Mixed Korean/English and table-spanning notes; spelling/wording needs review. |
| clinical trial summary | 75, 24+51, 13, 10, 52, 69%, 31 cases, 21 cases, 164 | Counts and percentages are small table text. |
| Masten/MDSGene | all cDNA/protein labels, counts 96, 74, 65, 64, 64, 55, 35, 543, and total 996 | Small figure labels. |
p.R275W total note | p.R275W: 54/996=6% | The chart label for p.Arg275Trp reads 64, but the note says p.R275W: 54/996=6%; preserve both until reviewed. |
| Morais Portugal text | 18 different mutations, 67.5%, 42.5%, 62.5%, 17/40, 9 different deletions, exon 4, exons 3-6 | Percentages and mutation/deletion counts are small text. |
| 40-patient table | all patient ages, cDNA/protein labels, family-history and consanguinity Yes/No values | Dense table; only selected row fragments are transcribed. |