PRKN Clinical Characteristics And Pathogenic Mutation Evidence
Total, 197 LoF+D.miss carriers exist in AMP-PD cohort
(Lesage, 2020 #1751)
- From 1500 cases, they found 199 Parkin mutant-PD
- 199 cases carried either homozygous (n = 92) or compound heterozygous (n = 59) PRKN mutations
Table 2. Demographic And Clinical Characteristics
TABLE 2. Demographic and Clinical Characteristics of Patients with Parkinson’s Disease with Pathogenic PRKN Variants (PRKN-PD) and of Patients without Pathogenic Variants (PD-NM)
| Characteristics | PD-NM n=1181 | PRKN-PD n=228 | p Value | Coefficient or odds ratio (OR) (SE) | Cohen's f2 | p Value |
|---|---|---|---|---|---|---|
| Baseline | ||||||
| Sex (% male) | 727/1,181 (61.6%) | 118/228 (51.8%) | 0.016 | |||
| Mean age at examination (SD), y | 50.5 (13.2) | 45.4 (12.9) | <0.0001 | |||
| Mean disease duration (SD), y | 8.8 (7.8) | 14.1 (10.4) | <0.0001 | |||
| Mean age at onset (SD), y | 41.6 (12.0) | 31.2 (10.7) | <0.0001 | |||
| L-DOPA-treated | 791/1,181 (67%) | 148/228 (64.9%) | 0.69 | |||
| Levodopa responsiveness | 659/738 (89.3%) | 142/149 (95.3%) | 0.045 | 1.9 (0.82) | 0.007 | 0.15 |
| Motor symptoms and signs | ||||||
| Dystonia at onset | 164/990 (16.6%) | 37/201 (18.4%) | 0.69 | 0.84 (0.18) | 0.001 | 0.46 |
| Akinesia at onset | 590/946 (61.3%) | 99/206 (48.1%) | 0.0010 | 0.51 (0.09) | 0.015 | 0.0003 |
| Tremor at onset | 570/960 (59.4%) | 142/205 (69.3%) | 0.021 | 1.7 (0.29) | 0.007 | 0.0076 |
| Micrographia | 308/927 (33.2%) | 42/204 (20.6%) | 0.0013 | 0.58 (0.11) | 0.007 | 0.010 |
| Asymmetry | 1,001/1,035 (96.7%) | 181/198 (91.4%) | 0.0049 | 0.26 (0.09) | 0.010 | 0.0005 |
| Bradykinesia | 1,033/1,069 (96.6%) | 201/208 (96.6%) | 1.0000 | 1.4 (0.63) | 0.002 | 0.46 |
| Rigidity | 1,008/1,066 (94.6%) | 194/204 (95.1%) | 0.90 | 1.1 (0.42) | <0.001 | 0.77 |
| Tremor | 796/1,056 (75.4%) | 168/205 (82%) | 0.057 | 1.5 (0.32) | 0.002 | 0.072 |
| Mean UPDRS III "ON" state (/108) (SD) | 19.6 (13.8) | 15.9 (11.9) | 0.0049 | -3.3 (1.2) | 0.008 | 0.014 |
| Mean Hoehn & Yahr "ON" state (/5) (SD) | 2 (0.91) | 2.00 (0.93) | 0.74 | -0.14 (0.09) | 0.003 | 0.16 |
| Dyskinesia | 457/667 (68.5%) | 97/178 (54.5%) | 0.0025 | 0.44 (0.09) | 0.020 | 0.0005 |
| Motor fluctuations | 485/663 (73.2%) | 82/177 (46.3%) | <0.0001 | 0.32 (0.07) | 0.041 | <0.0001 |
| Non-motor symptoms and signs | ||||||
| Dysautonomia | 254/470 (54.0%) | 36/192 (18.8%) | <0.0001 | 0.19 (0.05) | 0.095 | <0.0001 |
| Dementia | 67/667 (10.0%) | 6/153 (3.9%) | 0.037 | 0.34 (0.16) | 0.009 | 0.014 |
Data are expressed as mean (standard deviation) for continuous variables, and as counts (percentages) for categorical variables. We used t-tests to compare the two groups for continuous variables and Fisher’s exact tests for binary variables. Coefficients for continuous clinical features and odds ratios (ORs) for binary clinical features and standard error (SE), Cohen’s f2 and p-values were calculated from GLMs with mutation status, sex, age at onset, disease duration, L-DOPA group and age at onset vs disease duration for all 15 variables except for onset variables for which only mutation status, sex and age at onset were added. Linear models were used for continuous variables; GLMs with logit links and Bernoulli distributions were used for binary variables; UPDRS III, the motor subsection of the Unified Parkinson’s Disease Rating Scale.
Levodopa responsiveness was defined as a >30% improvement in subjective perceived motor symptoms.
PRKN Exon / Domain Variant Map
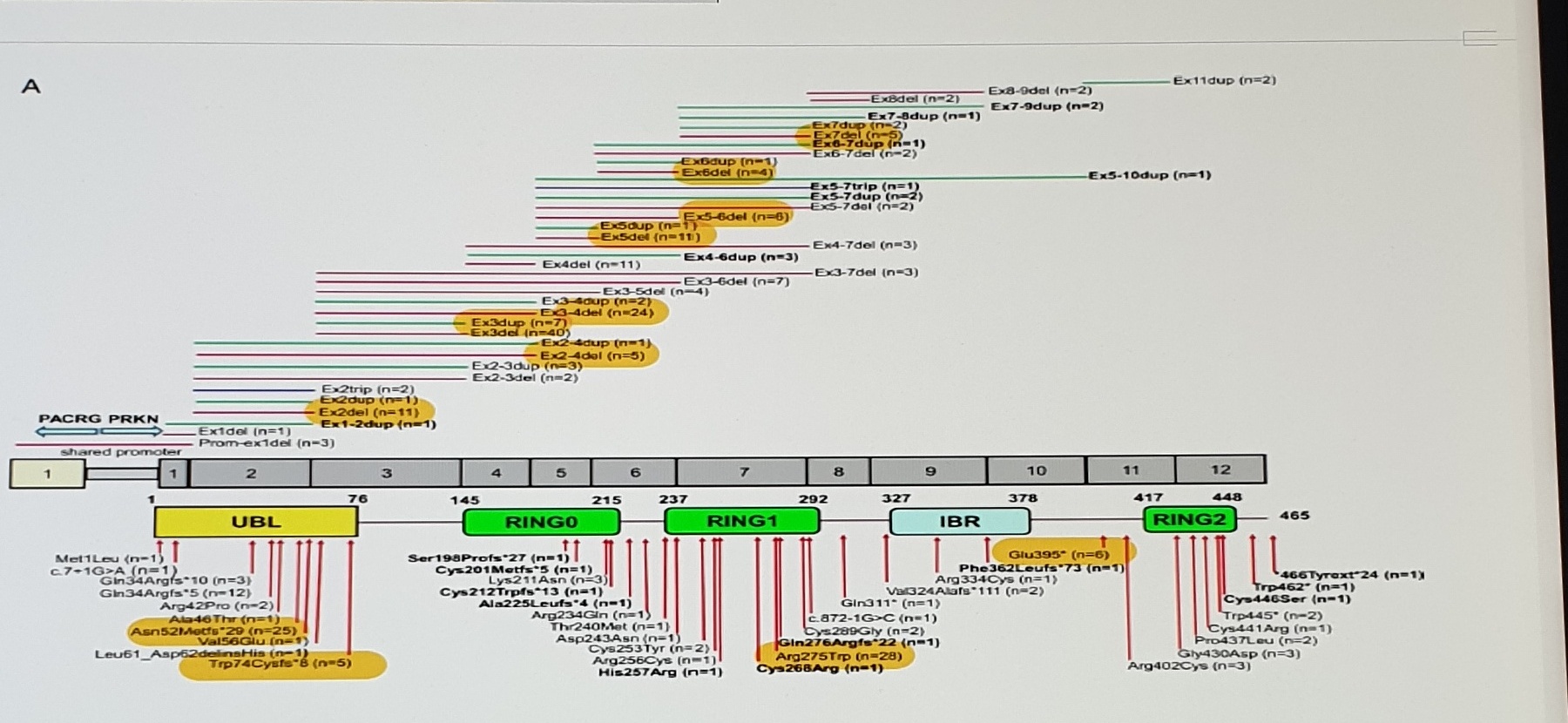
[Uncertain]ski, 2012 #692
the authors did two analysis at the same time
- UK Cohort 136명의 EOPD (이중 29 가 familial cases) 뒤져서 각각 다른 5개의 아래의 pathogenic Parkin mutation발견함 (모두 compound heteroz임, 그래서 아래에 mutation 1 & 2보임). (js: mean age: 52),
TABLE 2. Pathogenic mutations identified with descriptive clinical
| A | B | C | D | E | |
|---|---|---|---|---|---|
| Gene Mutation 1 | PARK2 42P | PARK2 c.154delA frameshift | PARK2 Del X2-3 | PARK2 c.438del40 frameshift | PARK2 Dup X3 |
| Mutation 2 | Dup. X3 | R275W | Del. X2 | Del. X2 | Dup X7 |
| Age | 49 | 58 | 49 | 53 | 50 |
| AAO | 37 | 35 | 8 | 20 | 20 |
| Ethnicity | Welsh | Welsh | Welsh | British | British |
| Sex | Male | Male | Male | Female | Male |
| Family history | None | None | 1 Sibling | None | None |
| Onset symptom | Lower limb stiffness/rigidity | Symmetrical resting tremor | Lower limb stiffness/rigidity | Lower limb dystonia | Lower limb stiffness/rigidity |
*Parents unaffected. AAO, age at onset
More detailed clinical descriptions can be found in the supplementary material.
Systematic Review / ROPAD Notes
- Systematic review, 43 studies with 3952 (EOPD 일것) patients
- 이중 8.6%가 Parkin mutation (아래기술 보면 이들이 모두 biallelic) (95% CI, 6.0%-12.4%; Fig.1, Table S3).
- 44.1% homozygous, 55.9% compound heterozygous
- Proportion (the author says this is ‘pathogenic parkin mutation’)
- 15.5% in Familial EOPD vs 4.3% in sporadic EOPD
ROPAD
Skrahina, 2021 #1926): 10/1288 were biallelic prkn mutation. So this gives ~54,000 patients WW.
#682: only missense mutations!
I can’t find the mean age of the subjects in the paper.
PRKN Variant Annotation Table, Partial
| Index | Databases | Parkin Variant | Annotation with clinical evidence | Functional group | Annotation with clinical and functional evidence |
|---|---|---|---|---|---|
| Disease and population | p.R42P | Pathogenic | 1 | Pathogenic | |
| Disease and population | p.V56E | Pathogenic | 1 | Pathogenic | |
| Disease and population | p.K211N | Pathogenic | 2 | Pathogenic | |
| Disease and population | p.C212Y | Pathogenic | 1 | Pathogenic | |
| Disease and population | p.C238W | Pathogenic | 1 | Pathogenic | |
| Disease | p.C253Y | Pathogenic | 1 | Pathogenic | |
| Disease and population | p.G284R | Pathogenic | 2 | Pathogenic | |
| Disease and population | p.T415N | Pathogenic | 2 | Pathogenic | |
| Disease | p.C431F | Pathogenic | 2 | Pathogenic | |
| Disease and population | p.C441R | Pathogenic | 1 | Pathogenic | |
| Disease | p.T240R | Likely Pathogenic | 2 | Pathogenic |
Uncertain Spans
- The citation prefix before “ski, 2012 #692” is cropped off at the left edge.
- Superscript footnote letters in Table 2 are not consistently legible; numeric values are retained without superscript letters where uncertain.
- Dense mutation labels inside the PRKN exon/domain map are preserved as an image rather than fully retyped because several labels are too small for lossless text reconstruction.
- The PRKN variant annotation table continues below the crop; only visible rows are transcribed here.